04-May-2020
Undeterred by the extraordinary circumstances created by the coronavirus pandemic, Instruct ES and INB/ELIXIR-ES scientists have been working hard to deliver tools and resources to support essential COVID-19 research.
On 4 May 2020, a team at the Instruct Image Processing Centre (hosted at the Centro Nacional de Biotecnología, CNB-CSIC in Spain) together with INB/ELIXIR-ES launched 3DBionotes-Covid19, a specialised, web-based application for the visualisation and annotation of protein structures relating to SARS-CoV-2. 3DBionotes-Covid19 has been developed from the 3DBionotes platform, an ELIXIR Recommended Interoperability Resource (RIR) that integrates protein structure, sequence and annotations through an interactive graphical interface (Fig 1).
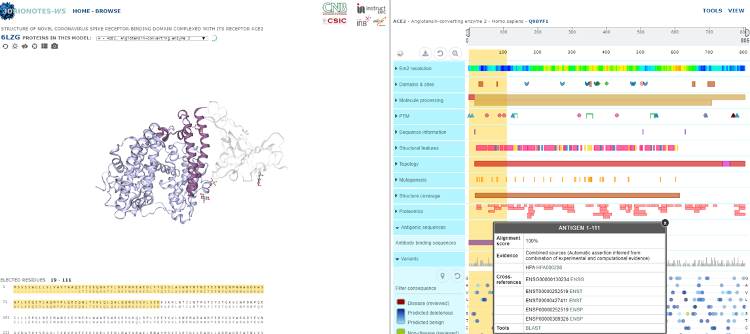
Fig 1. A screenshot from the 3DBionotes-Covid19 application showing the structure of the novel coronavirus spike receptor-binding domain complexed with its receptor ACE2 (left) and associated annotations, including sequence variants (right).
3DBionotes-Covid19 contains SARS-CoV-2 protein models from a variety of sources, including PDB-Redo and SwissProt, making special emphasis on the usage of the repository of the Coronavirus Structural Taskforce maintained by Prof. Andrea Thorn group. Structure models can be extensively annotated with on-protein and genomic annotations from the UniProt databases, as well as custom-made functional assignments from related sequences. Also, through a long-standing collaboration with Dr Ardan Patwardhan and Dr Sameer Velankar of EMBL-EBI, 3DBionotes-Covid19 provides annotations of quality-related parameters for cryoEM structures, such as local resolution and Q-score. And to help in the analysis of potential antiviral targets, 3DBionotes-Covid19 also supports the visualisation of interacting partners, such as drugs, and ligands from fragment-based screening experiments (including an XChem screen at the Diamond Light Source (Instruct Centre UK), to be followed by other related computational and functional efforts).
3DBionotes-Covid19 is hosted by the Biocomputing Unit of the CNB-CSIC in Madrid under the leadership of Prof Jose Maria Carazo and Dr Carlos Oscar Sanchez Sorzano, and is delivered in cooperation with Instruct-ERIC and ELIXIR through the Instruct Image Processing Centre (I2PC) and the INB/ELIXIR-ES node.
On 8 July 2020, an update was made to the 3DBionotes-COVID-19 website in order to collect all available SARS-CoV-2 structural data.
The update offers some very interesting new features, as well as some changes that are not evident on the ‘outside’ but are part of a huge effort to improve performance, consolidate data and refactor code.
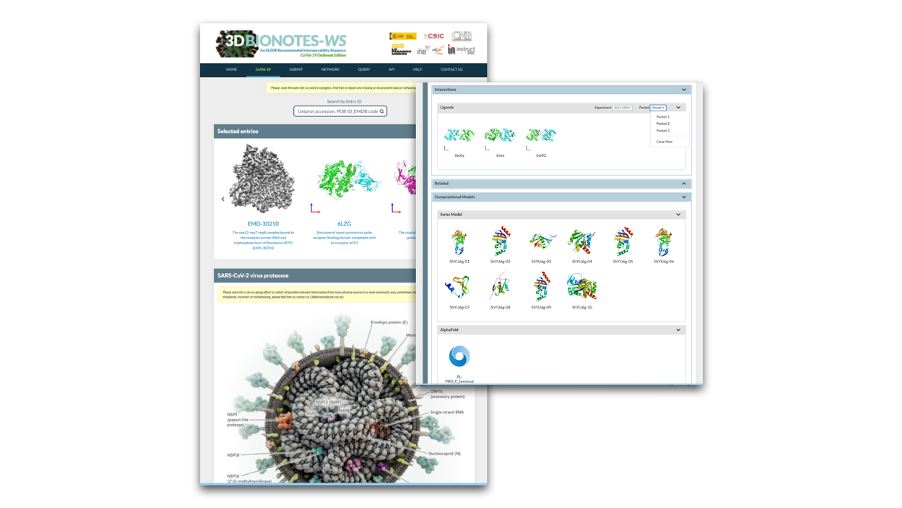
Fig 2. 3DBionotes includes a new way to move around by clicking directly on the label of the protein of interest in the new hyper realistic infographics. Also, the Ligands Interactions panel features new filters for the type of screening/repurposing experiment and binding pocket.
New features include:
Updates include:
---
Several digital initiatives have been delivered by Instruct scientists in response to the COVID-19 pandemic, including the crowdsourced COVID-19 page on Proteopedia: the free, 3D-encyclopedia of proteins and biomolecules, founded by Profs. Jaime Prilusky and Joel L. Sussman at the Weizmann Institute of Science (Instruct Centre IL). The Proteopedia COVID-19 page contains a variety of resources that have been created and curated by scientists from around the world, to present the latest findings in structural biology in a widely accessible way (Fig 3).

Fig 3. Proteopedia contains a variety of resources relating to COVID-19.