20-Mar-2020
To ensure cell survival, growth and differentiation, gene expression must be continuously modulated. Not all gene products are useful at every stage of cell life and the bulk of the genome is silent, packaged as chromatin (DNA wrapped around histone proteins), which limits access to the regulatory DNA sequences upstream of a gene. SAGA (Spt-Ada-Gcn5-Acetyltransferase) is a multi-protein complex involved in cell differentiation and the expression of protein-encoding genes. To understand how SAGA regulates gene transcription, scientists working at Instruct Centre FR1 at the IGBMC in Strasbourg have used national cryo-electron microscopy (cryo-EM) facilities to determine the atomic structure of yeast SAGA to 3.5 Å, which they published in Nature.
SAGA subunits
As a co-activator, SAGA (which comprises 19-subunits organised into four modules) is involved in a network of interactions that regulate mRNA synthesis by modifying the structure of chromatin in response to cell signals. One of SAGA’s roles is to modify chromatin structure through two enzymatic activities: acetylation and deubquitination. For example, the Gcn5 subunit of SAGA can catalyse the acetylation of specific lysine residues of histone proteins, which causes chromatin to decompact. SAGA also interacts with the TATA-box binding protein (TBP), which is crucial for the transcription of all genes as it initiates the assembly of the transcription machinery. More specifically, SAGA plays a key role in regulating the precise binding of TBP to the gene promoter and prevents the illegitimate transcription of non-coding DNA sequences.
Structural information about SAGA subunits
Using cryo-EM, the scientists investigated the architecture of SAGA when bound to TBP, since spatial organisation provides information about biological functions and the mechanisms that regulate the positioning of TBP. Cryo-EM showed that one of the two lobes that constitute SAGA is almost entirely occupied by the giant protein subunit Tra1, which is formed of 3800 residues organised in helical-loop-helical repeats (known as HEAT) and an inactive kinase domain (Fig 1). The Tra1 module is essential for interaction with the activating proteins that recruit SAGA upstream of the gene to be transcribed.
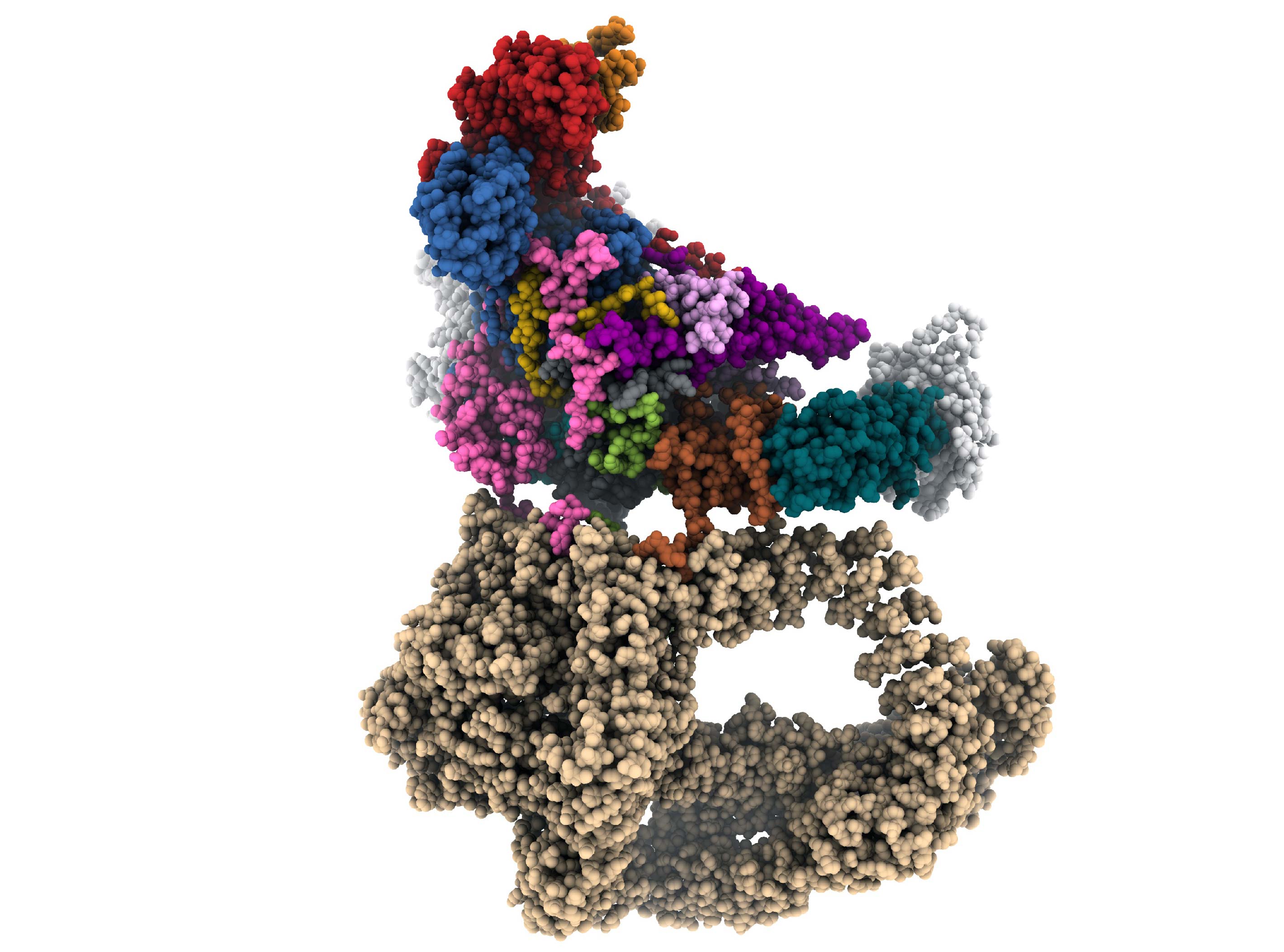
Fig 1. Front view of a composite cryo-EM reconstruction of the yeast SAGA–TBP complex. Maps from focused refinements of Tra1 (tan), the Spt8 (white), TBP (blue-green) and the main lobe were combined. HAT and DUB density are at lower resolution due to the flexibility of these modules and are not represented.
Cryo-EM also characterised several protein bridges that provide communication between Tra1 and all other SAGA subunits, which are located in the main lobe. These bridges are vectors for allosteric regulation between enzyme activities and transcriptional activator binding. The enzyme modules that allow chromatin nucleosome modification through acetylation and de-ubiquitination are highly mobile and are attached to the SAGA core by flexible protein bonds. It is proposed that the freedom of movement of the enzyme modules is necessary to recognise the substrate, once SAGA has bound to DNA via activators. The main lobe of SAGA is dominated by the Taf5 subunit, which plays a central organising role. Eleven protein domains of SAGA contact the WD40 domain of Taf5, making it a hub for connection and communication within the SAGA assembly.
TBP binding and structural changes within SAGA
It had previously been proposed that SAGA contains a protein octamer, similar in structure to the nucleosomal histone protein. Using cryo-EM, Papai et al. have been able to present an atomic structure of the full histone-fold octamer at the core of SAGA, which was crucially found to be asymmetric as compared to the nucleosomal histones. The authors propose that the octameric structure in SAGA is distorted to allow the TBP-binding subunit, Spt3, to interact with TBP.
The distortion of the histone-fold was found to be due to a 20-degree tilt of the Spt3 histone fold pair. Significantly, this tilt contributes to control the interaction between TBP and DNA. The distorted histone octamer forms a docking site that sequesters TBP in a conformation which prevents it from binding to DNA. In the presence of the general transcription factor TFIIA, TBP is released from SAGA, allowing it to interact with DNA to form a productive TBP-TFIIA-DNA complex. The release of TBP from SAGA can now be understood as being essential for the subsequent steps in assembling the transcription pre-initiation complex (PIC). In this way, the structure of SAGA regulates TBP's access to the promoter sequences of the genes, and hence transcription.
Thus, cryo-EM has provided new insights into the multiple roles of SAGA in regulating gene transcription, particularly the role of its protein subunits and the regulation of TBP function.
Find out more about the cryo-EM facilities that are available through Instruct-ERIC.
Read the paper: Papai, G., et al. (2020) Structure of SAGA and mechanism of TBP deposition on gene promoters. Nature 577, 711–716.