Instruct Centre ES launches 3DBionotes-Covid19: an online tool to study coronavirus proteins
On 4 May 2020, a team at the Instruct Image Processing Centre (hosted at the Centro Nacional de Biotecnología in Spain) together with INB/ELIXIR-ES launched 3DBionotes-Covid19, a specialised, web-based application to visualise and annotate protein structures related to SARS-CoV-2. 3DBionotes-Covid19 was developed from the 3DBionotes platform, an ELIXIR Recommended Interoperability Resource that integrates protein structure, sequence and annotations through an interactive graphical interface.

3DBionotes-Covid19 is an online platform to study coronavirus proteins in three dimensions.
As the scientific community have responded to the COVID-19 pandemic, a variety of experimental tools, including cryo-EM, X-ray crystallography and NMR, have been used to obtain detailed atomic models of the SARS-CoV-2 proteins in different conformations and complexed with different partners (including the main receptor in human cells). To make this important structural information more accessible, data for SARS-CoV-2 proteins in their genomic, proteomic and interatomic context have been made available via the 3DBionotes-Covid19 platform.
Besides structural data, 3DBionotes-Covid19 also contains a wealth of scientifically interesting information, including the local quality of the experimental data (so that the structure in that region can be more or less trusted); molecular patterns (domains and folds) known to have particular roles or interactions with other molecules; genetic variants of the virus observed in COVID-19 patients; molecular modifications (eg glycosylation) known to occur in those regions; and interactions with small drug fragments or drug interaction predictions. This information is helping scientists to better understand SARS-CoV-2 viral proteins and their role in infection, and to identify possible targets for therapeutics or preventive treatments.
On 8 July 2020, an update to 3DBionotes-COVID-19 introduced some interesting new features, as well as upgrades to improve performance, consolidate data, and refactor code.
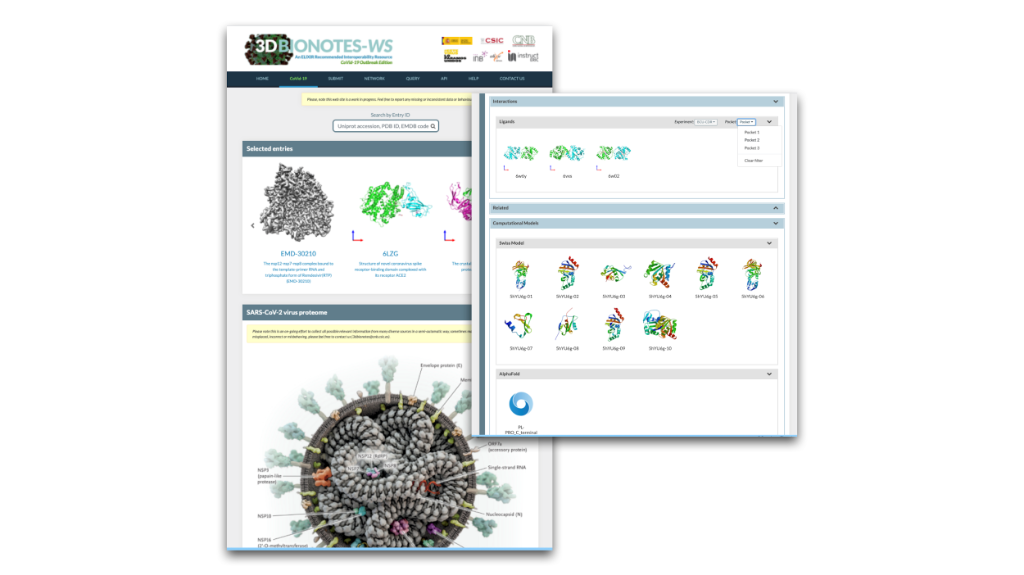
3DBionotes includes a new way to move around by clicking directly on the label of the protein of interest in the new hyper realistic infographics. Also, the Ligands Interactions panel features new filters for the type of screening/repurposing experiment and binding pocket.
New features include:
- hyper realistic infographics representing a virus used as a map index. Users can click on a protein label and will be directed to the corresponding section
- additional computational models from various sources: SwissModel, AlphaFold & BSM-Arc. These are especially useful for those proteins with missing experimental models
- a ligand interactions section, including results from experimental and computational drug screening & repurposing (one from the I2PC group is to be published in short)
- filters for ligand interactions, enabling selection by experiment and interaction pocket.
Updates include:
- Improved loading for validated models from PDB-Redo & Isolde
- Updated help & documentation.
3DBionotes-Covid19 is hosted by the Biocomputing Unit of the CNB-CSIC in Madrid under the leadership of Prof Jose Maria Carazo and Dr Carlos Oscar Sanchez Sorzano, and is delivered in cooperation with Instruct-ERIC and ELIXIR through the Instruct Image Processing Centre (I2PC) and the INB/ELIXIR-ES node.
Key structures
| EMPIAR-10516 | Cryo electron microscopy of SARS-CoV-2 spike in prefusion state | 2020-09-22 |
| EMD-11336 | SARS-CoV-2 spike in prefusion state (flexibility analysis, 1-up closed conformation) | 2020-07-29 |
| EMD-11337 | SARS-CoV-2 spike in prefusion state (flexibility analysis, 1-up open conformation) | 2020-07-29 |
| EMD-11338 | SARS-CoV-2 spike in prefusion state | 2020-07-29 |
| EMD-11341 | SARS-CoV-2 stabilized spike in prefusion state (1-up conformation) | 2020-07-29 |
| 6ZP5 | SARS-CoV-2 spike in prefusion state (flexibility analysis, 1-up closed conformation) | 2020-07-29 |
| 6ZP7 | SARS-CoV-2 spike in prefusion state (flexibility analysis, 1-up open conformation) | 2020-07-29 |
| 6ZOW | SARS-CoV-2 spike in prefusion state | 2020-07-29 |
Visit the 3DBionotes-Covid19 website to find out more.
